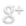Team Advisors
Andrew Mendelsohn
Jennifer Lei,
Team Members
Gabriel Sumano, UC Berkeley ’09
Ida Bezabeh, UCLA ’11
Israel Lipinksky, UC Berkeley ’08
Justine Widjaja, UC Berkeley ’11
Peijia Yuan, UC Berkeley ’12
Tarang Srivastava, UC Berkeley ’11
The Project
Breast cancer is the second most common type of cancer diagnosed in women. More than 40,000 women are expected to die this year from breast cancer. Mortality results primarily from complications of metastatic breast cancer. New therapeutics are desperately needed. We have designed a novel system to target malignant breast cancer cells. We based our system on the pioneering work of Hajitou and colleagues, who built a single-stranded DNA bacteriophage/Adeno Associated virus (AAV) hybrid that delivers DNA encoding tumor-killing proteins to cancer cells. We extended their system by incorporating a module to express RNAi knock down expression of critical oncogenes. Furthermore, to solve the problem of strong neutralizing immune responses against bacteriophage in humans, we designed our hybrid vector to bind sponge-derived siliconizing enzymes that can encapsulate the hybrid virus in silicates. This not only should attenuate any immune response, but also opens the door to create chimeric cyborg-like nanomachines. For proof of principle, we inserted DNA encoding a small piece of the Raf-1 proto-oncogene into an AAV cassette containing two Inverted Terminal Repeats (ITR’s) that help efficiently transduce mammalian cells. This cassette was then inserted into fd-derived bacteriophage vector, FUSE55. To target the malignant subset of breast cancer cells that express Her2, DNA encoding a single chain antibody to Her2 was fused to DNA encoding gene III, a coat protein. The fusion protein also contained the Streptavidin II tag, which allows it to bind sponge-derived silicateins and silaphantin. We believe that our hybrid, siliconizing virus represents both a powerful new approach to cancer therapy and a promising model system to develop silicon/carbon nanomachines.
Motivation
Breast cancer is one of the more common kinds of cancer among women. We have designed a new treatment delivery method using Her2 receptors, which are over-expressed in about a quarter of breast cancers. Breast cancers that over-express Her2 have poorer prognosis than breast cancers without Her2 amplification. Therefore, we have attempted to build a viable system for the delivery of RNAi into breast cancer cells with many copies of the Her2 receptor.
Breast cancer is the seventh leading cause of death of women in the US, and the leading type of cancer diagnosed in women. To date, there are no therapies that specifically target breast cancer cells while delivering RNAi. Instead, current methodologies still rely on surgical excision of the malignant growth, non-specific systemic administration of cytotoxic drugs, or antibodies that induce non-specific apoptosis by an unknown mechanism. Therefore, to address the need for a novel breast cancer therapy, we designed a method of phage-mediated delivery of shRNA using a breast cancer-specific single-chain antibody that specifically targets breast cancer cells. In addition, to decrease the immunologic response to possible phagemid proteins, we encased the phage in a silica coat. We hope that our system opens the gateway for possible novel cancer therapies not premised on the classical technologies.
Experimental Overview
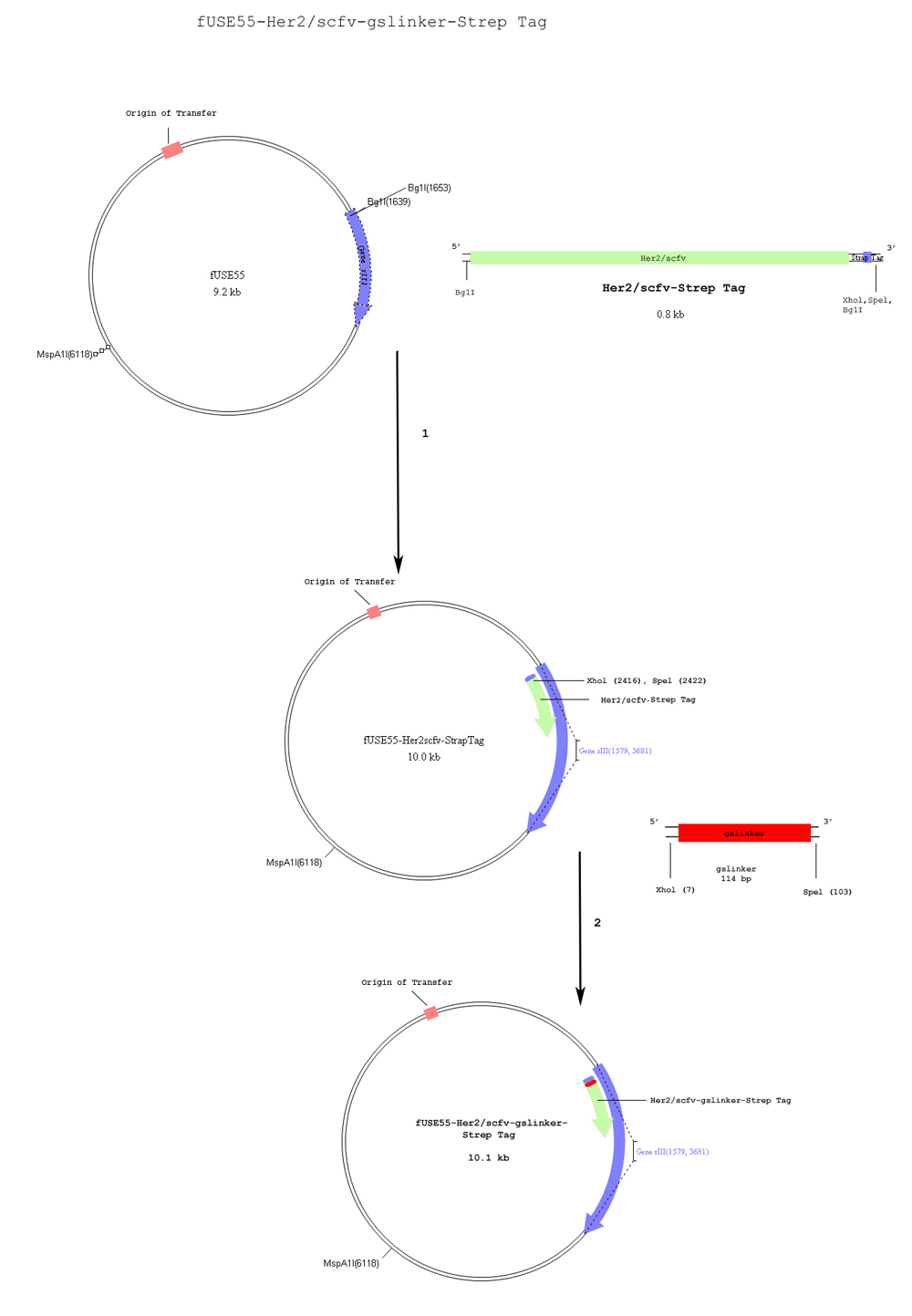
We targeted cells with Her2 amplification and used RNAi to turn off the Raf-1 gene, which is constitutively on in many cancers. First, we inserted an shRNA sequence coding for mRNA causing knockdown of Raf-1 expression into an AAV cassette containing two ITR’s, allowing for episomal replication. This cassette was inserted into our chosen vector, the filamentous bacteriophage FUSE55. An antibody sequence was then added to the phage plasmid in order to target Her2. As an additional feature, we have fused Silicatein and Silintaphin to mStreptavidin, which will bind to a protein tag in the coat. This will cause silicate structures to form on the coat of the phage, thereby reducing the immunotoxicity of the bacteriophage in vivo.
Project Details
Since the beginning of molecular biology, phage have been extensively used as a means to understand the transfer of genetic material. Additionally, owing to their evolutionary perfection for selfish replication, they have small genomes and are amenable to extensive genetic manipulation. One useful aspect of phage is the ability to display foreign proteins on the surface, such as in phage display libraries. Thus, we utilized phage as a platform for the targeting and introduction of small interfering RNA (RNAi) into breast cancer cells (1).
Phage Selection
Phage as a vehicle for gene delivery
Since the beginning of molecular biology, phage have been extensively used as a means to understand the transfer of genetic material. Additionally, owing to their evolutionary perfection for selfish replication, they have small genomes and are amenable to extensive genetic manipulation. One useful aspect of phage is the ability to display foreign proteins on the surface, such as in phage display libraries. Thus, we utilized phage as a platform for the targeting and introduction of small interfering RNA (RNAi) into breast cancer cells (1).
Phage selection
Initially, single-stranded RNA phages such as Q-beta and MS2 appeared to be viable options. The single-stranded form of RNA was crucial in forming the short hairpin structure. In addition, the project design required a phage with a small genome size that has the ability to display fusion proteins on its coat surface. MS2 appeared to be the best option; however, further research on this phage showed a lack of information on its history of effectiveness in vitro. Since RNA phages did not match our criteria, DNA phages presented themselves as the next logical option. Phage fd-tet has a single phage chromosome (genome) bearing a single gene III coat protein, which accepts foreign DNA inserts (3). However, only 3-5 pIII (protein encoding geneIII)are displayed on the phage surface. It was necessary for the phage to accept foreign DNA for the display of the Her2 antibody and the targeting of breast cancer cells(4). Our method of Phage fd-tet targeting has been successfully shown to work in the past(2,5).
Previous studies have shown the ability of recombinant adeno-associated virus and phage to transduce mammalian cells in vitro (7). AAV vectors are also amenable to introduce short hairpin RNA structures which induce the RNAi pathway(8).
Specific Targeting of Breast Cancer Cells via a Her2 single chain antibody (Her2-scfv)
One of the hallmarks of cancer is uncontrolled, aberrant cellular growth. In most cancer types, there is an absence of specific cancer markers. Instead, cancer cells exhibit upregulation or downregulation of endogenous proteins. Thus, specific targeting of cancer cells is problematic. However, tumor specific targeting has produced effective results in the past and has even been shown to produce tumor regression(9). Her2 is over-expressed in about one-fourth of all cases of breast cancer and is utilised as a target for current monoclonal antibody therapies such as Herceptin. When Her2scfv-bearing phage bind to Her2 receptors, the phage-Her2 receptor complex enters the cell via endocytosis. Once internalised, the aav-shRaf-1 cassette will undergo independent episomal replication and produce shRaf-1 RNAs.
Silicatein and silintaphin have been used by sponges that have silica-based structures(11) and have been tested to encapsulate other structures such as bacteria(12). Silica is known to reduce immunotoxicity in the human body. Silintaphin and silicatein work together to produce silica structures for biomineralization and encapsulation of biological structures. As a result, the silicanized products are capable of entering biological system without inducing an immune response(13). We used two isoforms of silicatein (alpha and beta) from the silicaneous sponge, Tethya aurantia, and an optimised synthetic silintaphin from the silicaneous sponge, Suberites domuncula. The silicatein-silintaphin-streptavidin constructs were inserted into alphapp8-4m5.3, a yeast expression plasmid engineered for protein secretion. The silicatein-silintaphin-streptavidin fusion proteins were purified on streptactin columns and co-incubated with Her2-scfv-stepTag phage to allow for the silicanizing proteins to bind to the phage surface.
Potential Pitfalls
1) Efficacy of the shRaf-1 to inhibit expression of Raf-1. Basal expression of Raf-1 after knock down may allow for the persistence of the cancer cell into a more malignant form. 2) Silicanization of phage to circumvent immune response. Because only 3-5 pIII are displayed on the surface of the phage, there may be irregular silica formation. Also, optimisation of the linker regions between the proteins may be needed to allow optimal silica seeding. 3) Non-specific targeting of cells via the Her2 receptor
Future Plans/Endeavors
Future extensions to the aim of this project could be made in the form of modification of the specific types of receptors or gene sequences targeted for a more effective knockdown of various types of cancer cells. Genes to consider might include sequences involved in cell proliferation, anti-apoptosis, mitotic checkpoints, and so forth. On the other hand, the applications of the RNAi phage delivery system could be tested in different types of cells attacked by various types of cancer. Finally, different materials used in coating of the phage could be utilized and tested for better immunotoxicity response. Some other materials to consider might include gold, which has been known to not cause a reaction of the skin, and could thus be useful in targeting carcinomas.
Background
HER2
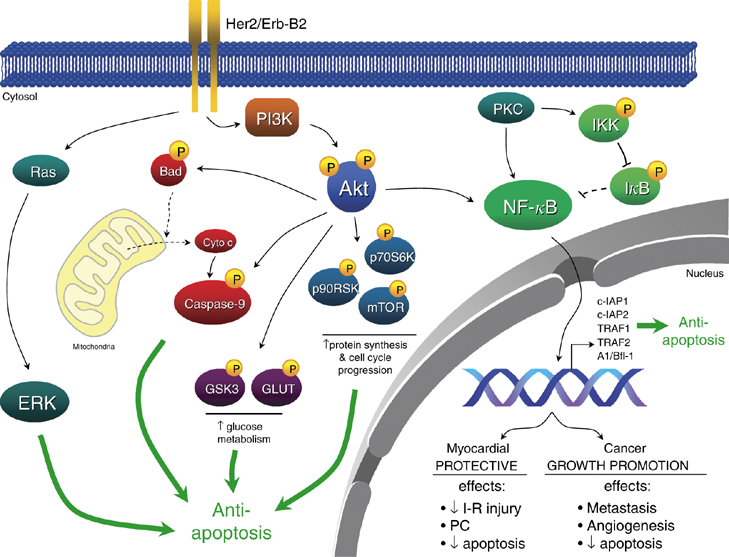
Her2 is a member of the erbB family of receptors and a receptor tyrosine kinase. ErbB receptors require a ligand such as a growth factor, and receptor tyrosine kinases add phosphate to tyrosine residues, essentially activating the receptor and allowing it to be a docking site for other proteins to become active. This receptor is involved in signal transduction mechanisms of cell proliferation. The erbB receptors require dimerization upon binding with its ligand EGF. Her2 itself has no ligand, but Her2 is the receptor that can dimerize with any of the other receptors in the family such as Her1 and Her3. Thus it is a favorite in terms of cancer because it is ligand-independent and quick to dimerize. It can become activated and provide a docking site for proteins to become active. One such protein affected by receptor tyrosine kinases is Raf-1.
RAF-1
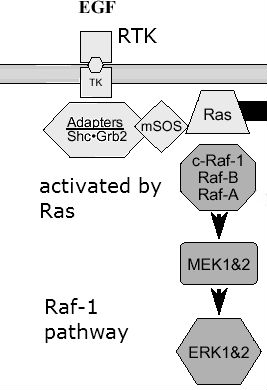
Raf-1 is a protein kinase involved in the MAP kinase pathway. Raf-1 is a MAP kinase kinase kinase which is activated by Ras. Once activated, it can phosphorylate downstream kinases like MEK1. Ultimately, this pathway helps in cell proliferation, differentiation, and migration, because MEK turns “on” ERK which is a transcription factor for genes such as cell proliferation.
Thanks to http://www.jpp.krakow.pl/journal/archive/1203_s3/articles/11_article.html:
Functional Designs Goals for Fuse55 Plasmid
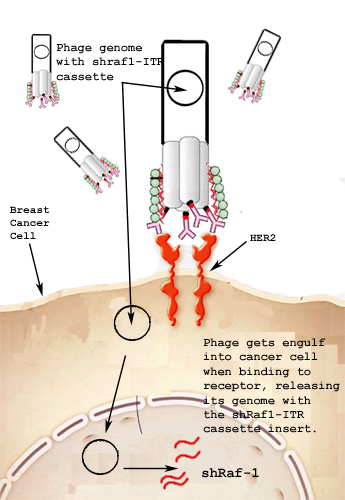
1. AAV vector is a part of the plasmid because it provides inverted terminal repeats (ITRs) which allow for the formation of hairpin structure due to its symmetry. It also assists in integration of viral DNA into genomic DNA of the host.
2. shRNA RAF-1 sequence forms the hairpin structure when it is transcribed. This allows it to be degraded in the RNAi process causing it to degrade other Raf-1 mRNAs, which match the sequence.
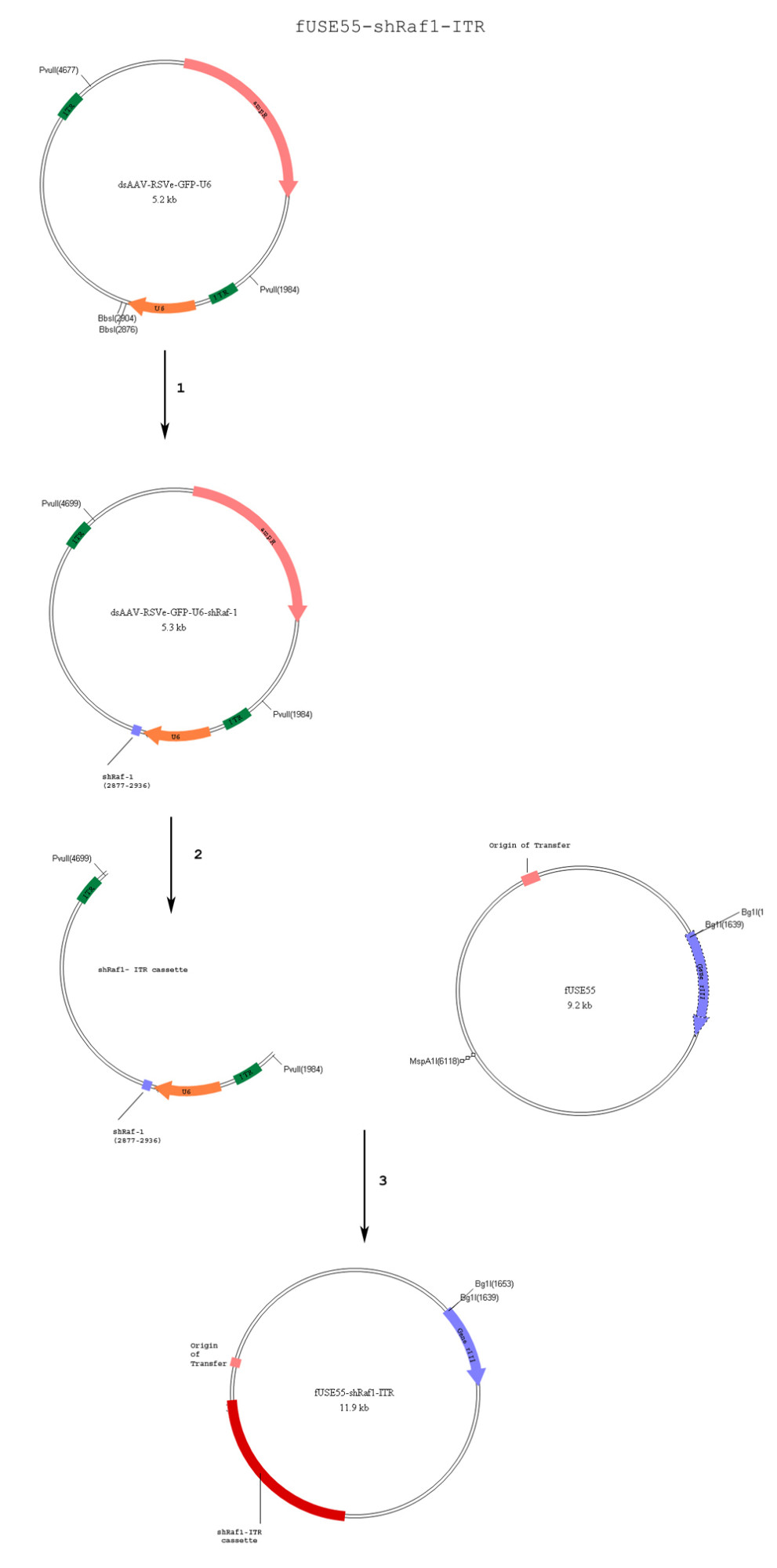
3. Her2 antibody binds to the Her2 receptor on breast cancer cells.
4. mStreptavidin to bind to silica globules and encapsulate the phage.
Results
References
1. Grimm, Dirk, and Mark A. Kay. “RNAi and Gene Therapy: A Mutual Attraction.” Hematology. (2007): 473-481.
2. Jiang, Hua, Xiu-mei Cai, Bi-zhi Shi, Jie Zhang, and Zong-hai Li. “Development of efficient RNA interference system using EGF-displaying phagemid particles.” Acta Pharmacol Sin. 29.4 (2008): 437-442.
3. “Explanation of the Smith Lab Phage Display Vectors” <www.biosci.missouri.edu/smithgp/PhageDisplayWebsite/vectors.doc>
4. Hajitou, Amin, Martin Trepel, et al. “A Hybrid Vector for Ligand-Directed Tumor Targeting and Molecular Imaging.” Cell. 125. (2006): 385-398.
5. Larocca, David, Paul D. Kassner, Alison Witte, Robert Charles Ladner, and Glenn F. Pierce. “Gene transfer to mammalian cells using genetically targeted filamentous bacteriophage.” FASEB. 13. (1999): 727-734.
6. Benhar, Itai, and Yoram Reiter. “Phage Display of Single-Chain Antibody Constructs.” Current Protocols in Immunology. (2002): 10.19B.1-10.19B.31.
7. Hajitou, Amin, et al . “Design and construction of targeted AAVP vectors for mammalian cell transduction.” Nature Protocols. 2.3 (2007): 523-531.
8. Grimm, Dirk, Kusum Pandey, and Mark A. Kay. “Adeno-Associated Virus Vectors for Short Hairpin RNA Expression.” Methods in Enzymology. 392. (2005): 381-405.
9. Eriksson, Fredrik, W. David Culp, Robert Massey, Lars Egevad, and Donita Garland. “Tumor specific phage particles promote tumor regression in a mouse melanoma model.” Cancer Immunology Immunother. 56. (2007): 677-687.
10. Sapinoro, Ramil, Ketna Volcy, W. W. Shanaka, I. Rodrigo, and Jacob J. Shlesinger. “Fc receptor-mediated, antibody-dependent enhancement of bacteriophage lambda-mediated gene transfer in mammalian cells.” Virology. 373.2 (2008): 274-286.
11. Muller, Werner, Xiaohong Wang, Fu-Zhai Cui, Klaus Peter Jochum, and Wolfgang Tremel. “Sponge spicules as blueprints for the biofabrication of inorganic–organic composites and biomaterials.” Appl Microbiology Biotechnology. 83. (2009): 397-413.
12. Muller, Werner, Sylvia Engel, Xiaohong Wang, Stephen Wolf, and Wolfgang Tremel. “Bioencapsulation of living bacteria (Escherichia coli) with poly(silicate) after transformation with silicatein-alpha gene.” Biomaterials. 29.7 (2008): 771-9.
13. Wiens, Matthias, Melanie Bausen, Felipe Natalio, Thorben Link, and Ute Schlossmacher. “The role of the silicatein-a interactor silintaphin-1 in biomimetic biomineralization.” Biomaterials. 30. (2009): 1648-1656.
14. Yarden, Yosef. “Biology of HER2 and Its Importance in Breast Cancer.” Oncology. 61. (2001): 1-13.